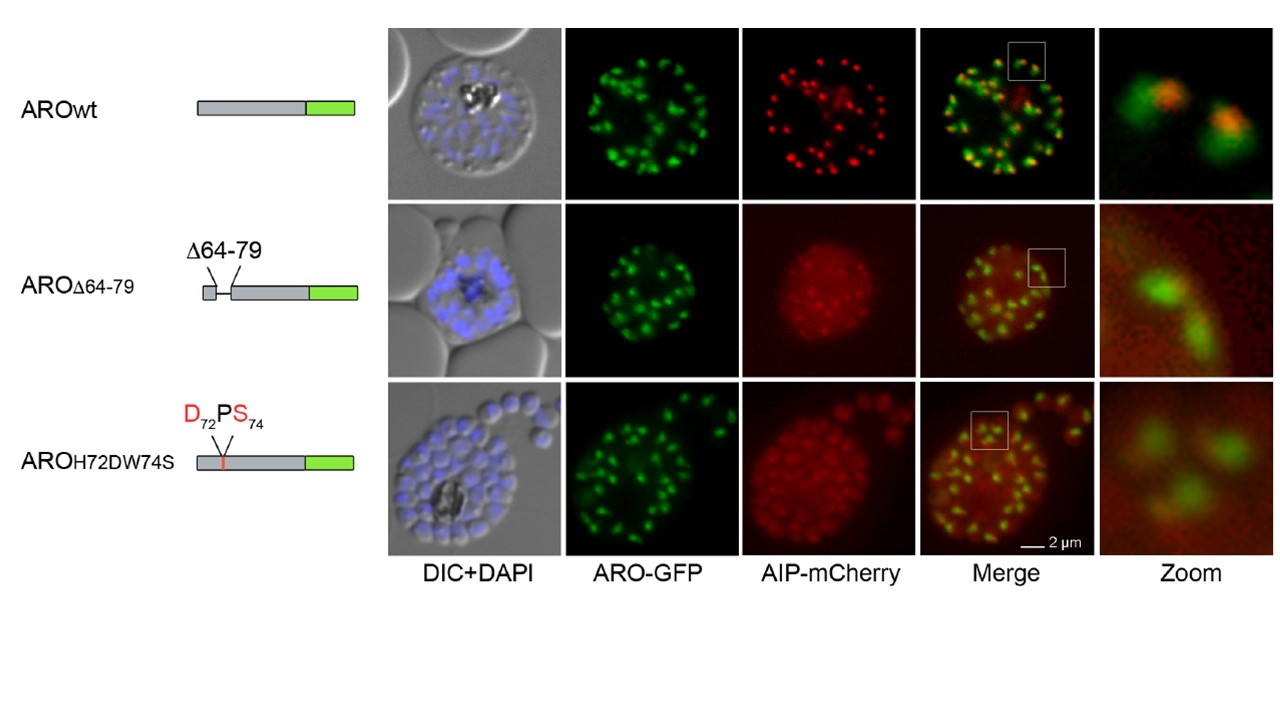
Mutations in putative PfARO interaction domain cause cytosolic distribution of PfAIP. (A) Schematics of the biscistronic expression plasmid pARL_PfARO-GFP_PfAIP-mCherry under the control of the AMA1 promoter. Grey PfARO coding region, green: GFP, blue: T2A skip peptide [64,65] turquoise: PfAIP coding sequence, red: mcherry (B) Schematic representation of the PfARO-GFP variants (PfAROWT, PfAROD64-79 PfAROH72DW74S) and their co-localization with PfAIPmCherry. Scale bars, 2 μm. Nuclei stained with DAPI. DIC, differential interference contrast; 5x zoom indicated by white square. While the co-expression of wild-type PfARO-GFP with PfAIP-mCherry leads to normal rhoptry association of both proteins (first row), the deletion of the loop region (AA 64-79) leads to re-distribution of PfAIP resulting in mostly cytosolic localization while the mutant PfARO is still targeted to the rhoptries (second row).
Geiger M, Brown C, Wichers JS, Strauss J, Lill A, Thuenauer R, Liffner B, Wilcke L, Lemcke S, Heincke D, Pazicky S, Bachmann A, Löw C, Wilson D, Filarsky M, Burda PC, Zhang K, Junop M, Gilberger TW. Structural insights into PfARO and characterization of its interaction with PfAIP. J Mol Biol. 2019 Dec 23. pii: S0022-2836(19)30733-8.
Other associated proteins
| PFID | Formal Annotation |
|---|---|
| PF3D7_1136700 | armadillo-interacting protein aip |