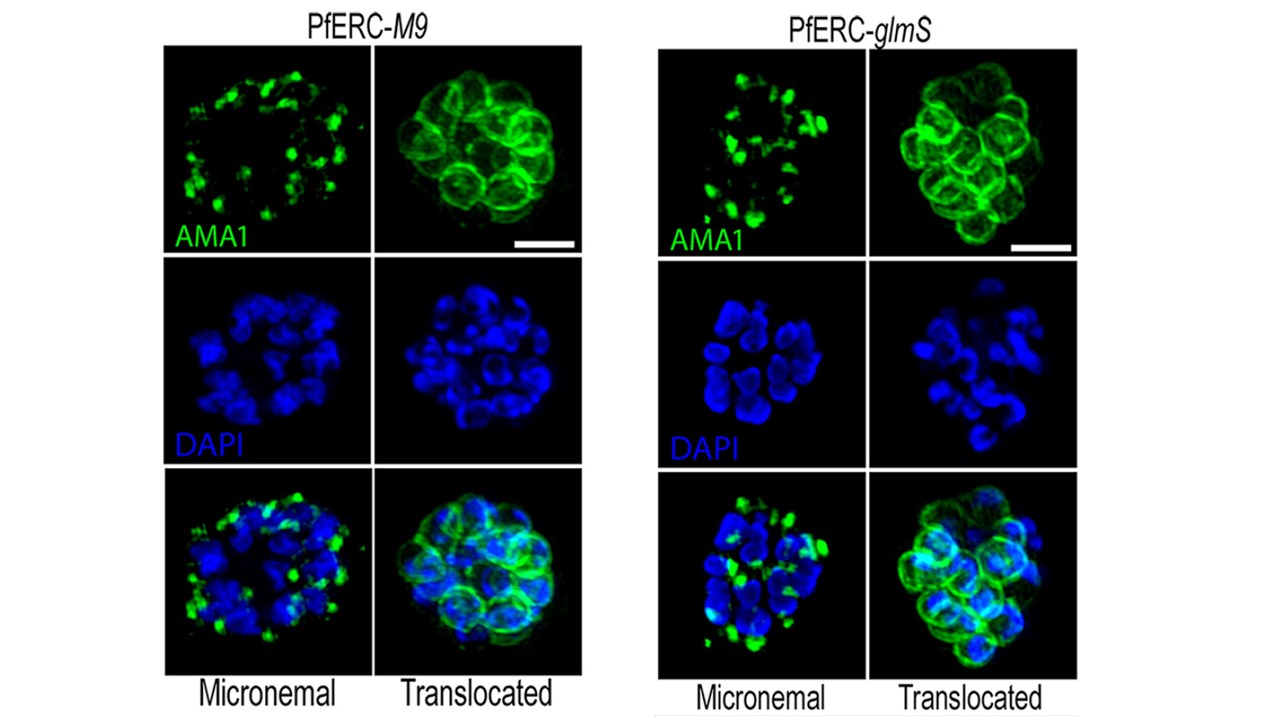
Proteolytic processing of AMA1 requires PfERC. (A) Parasites were grown in the presence of GlcN
for 48 h and then incubated with compound 1 for 4 h. Compound 1 was then removed, and parasites were incubated further with E-64 for 6 h and stained with anti-AMA1 as well as the nuclear stain DAPI.
Representative SIM images of these PfERC-glmS (PfERC tagged with the inducible version of the ribozyme gene, glmSn) and PfERC-M9 (inactive version of the ribozyme, M9) schizonts are shown. From top to bottom, the images are anti-AMA1 (green), DAPI (blue), and fluorescence merge. Bar, 2 mm. Using SIM, we observed AMA1 localization either in micronemes or on the surface of merozoites. Data show that there were no differences between the PfERC-M9 and PfERC-glmS parasites in the localization of AMA1, suggesting that PfERC does not function in the signaling pathway required for vesicle secretion.
Fierro MA, Asady B, Brooks CF, Cobb DW, Villegas A, Moreno SNJ, Muralidharan V. An Endoplasmic Reticulum CREC Family Protein Regulates the Egress Proteolytic Cascade in Malaria Parasites. mBio. 2020 Feb 25;11(1). pii: e03078-19.
Other associated proteins
| PFID | Formal Annotation |
|---|---|
| PF3D7_1108600 | endoplasmic reticulum-resident calcium binding protein |